Reconciling Membrane Protein Simulations with Experimental DEER

Award Winners and Abstracts of the 32nd Annual Symposium of The Protein Society; Boston, MA, July 9–12, 2018 - 2018 - Protein Science - Wiley Online Library

A new perspective on membrane-embedded Bax oligomers using DEER and bioresistant orthogonal spin labels

Full article: Membrane models for molecular simulations of peripheral membrane proteins

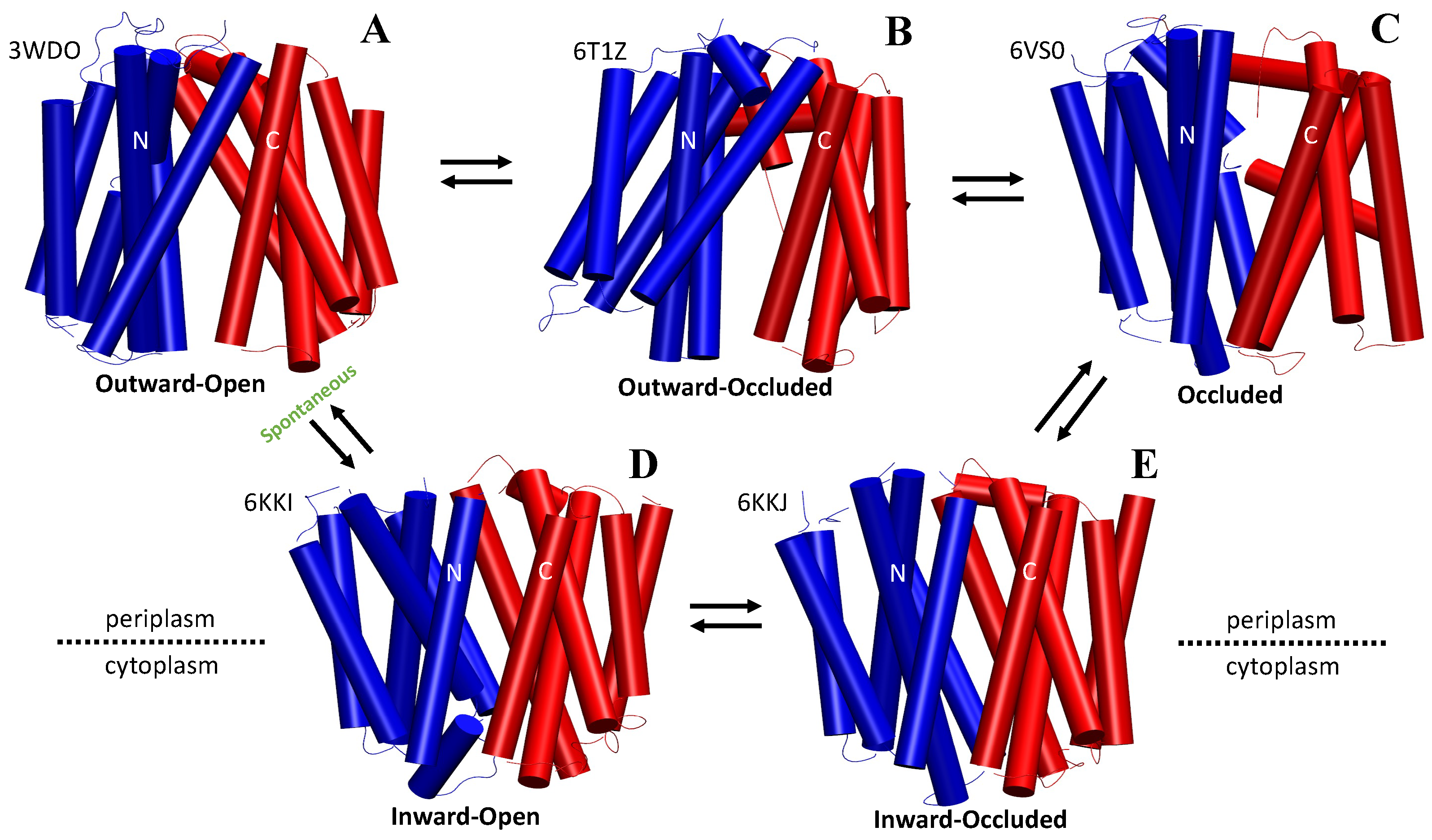
The ABC transporter MsbA adopts the wide inward-open conformation in E. coli cells
Membranes, Free Full-Text
Methodology for rigorous modeling of protein conformational changes by Rosetta using DEER distance restraints

DEER EPR Measurements for Membrane Protein Structures via Bifunctional Spin Labels and Lipodisq Nanoparticles

A) Cross-validated scores for 4300 possible DEER residue-pair sets.

DEER EPR Measurements for Membrane Protein Structures via Bifunctional Spin Labels and Lipodisq Nanoparticles

In situ observation of conformational dynamics and protein ligand–substrate interactions in outer-membrane proteins with DEER/PELDOR spectroscopy

Molecular dynamics simulations of biological membranes and membrane proteins using enhanced conformational sampling algorithms - ScienceDirect

PDF) Reconciling Membrane Protein Simulations with Experimental DEER Spectroscopy Data